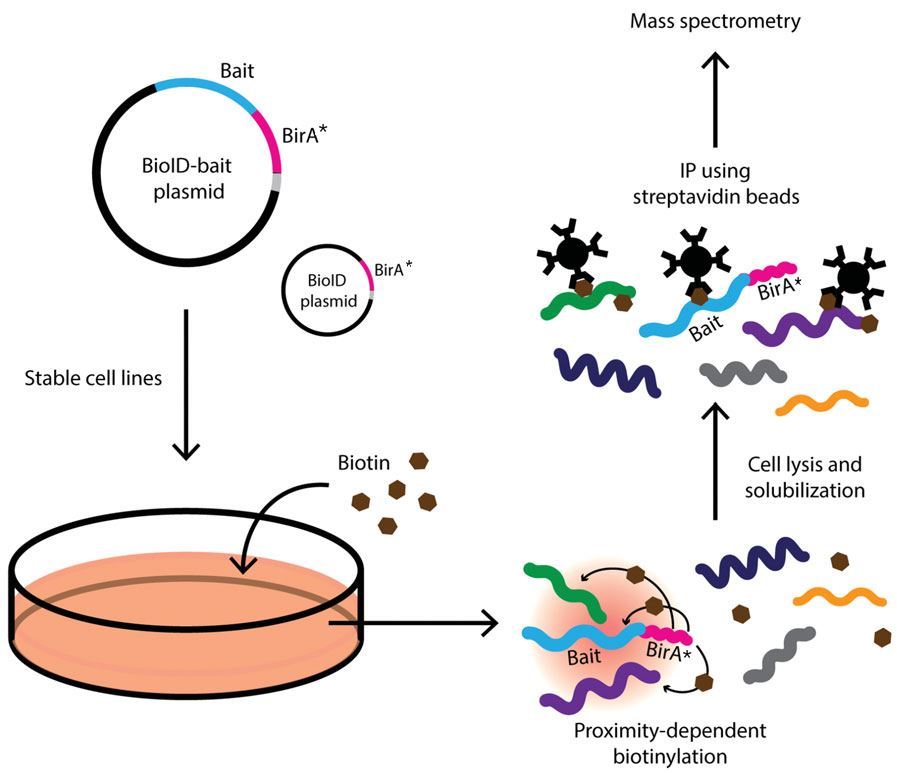
Identification of interacting proteins using proximity-dependent biotinylation with BioID2 in Trypanosoma brucei

In planta proximity-dependent biotin identification (BioID) identifies a TMV replication co-chaperone NbSGT1 in the vicinity of 126 kDa replicase - ScienceDirect

Probing mammalian centrosome structure using BioID proximity-dependent biotinylation - ScienceDirect
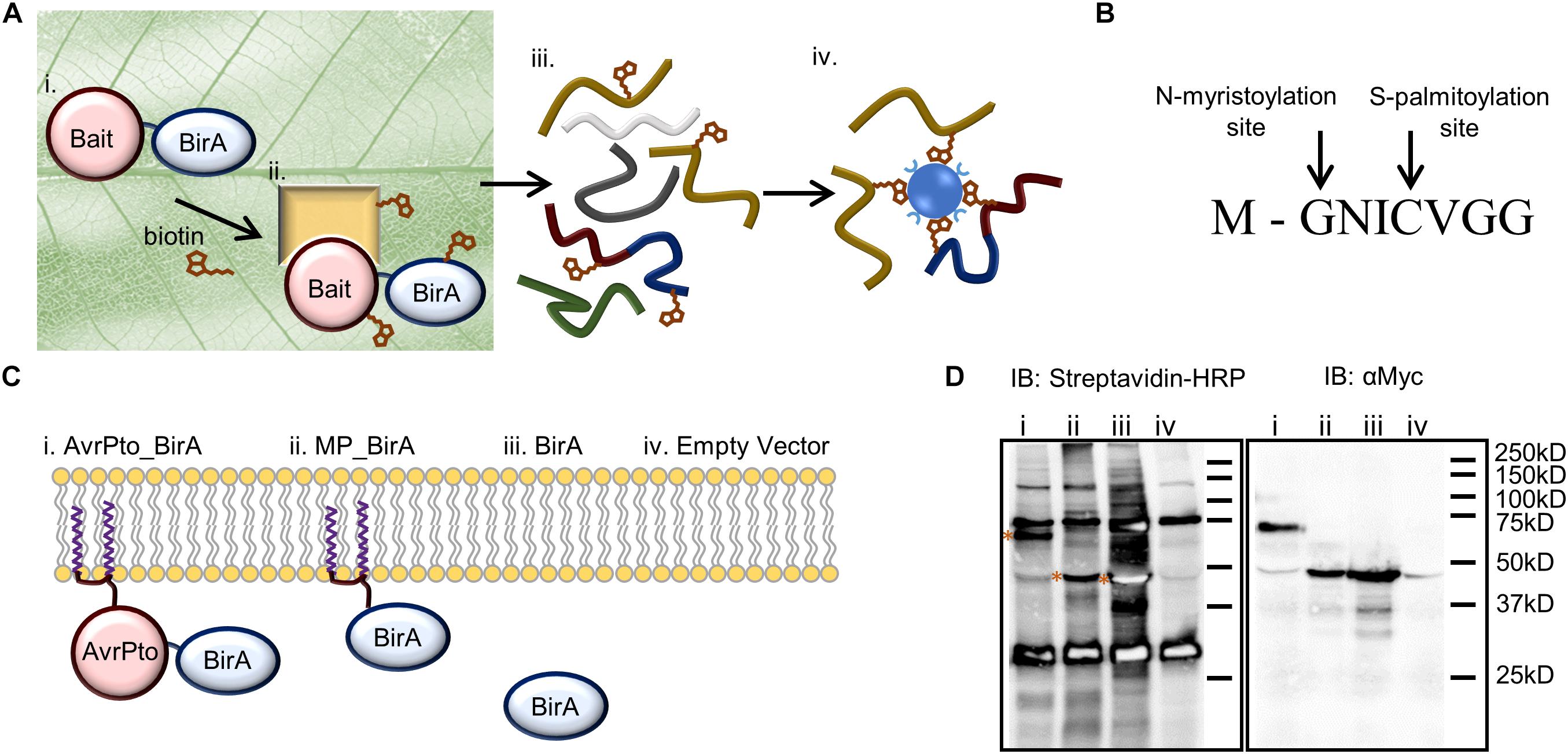
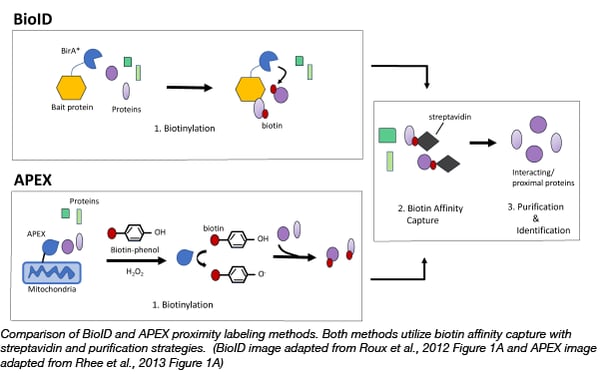
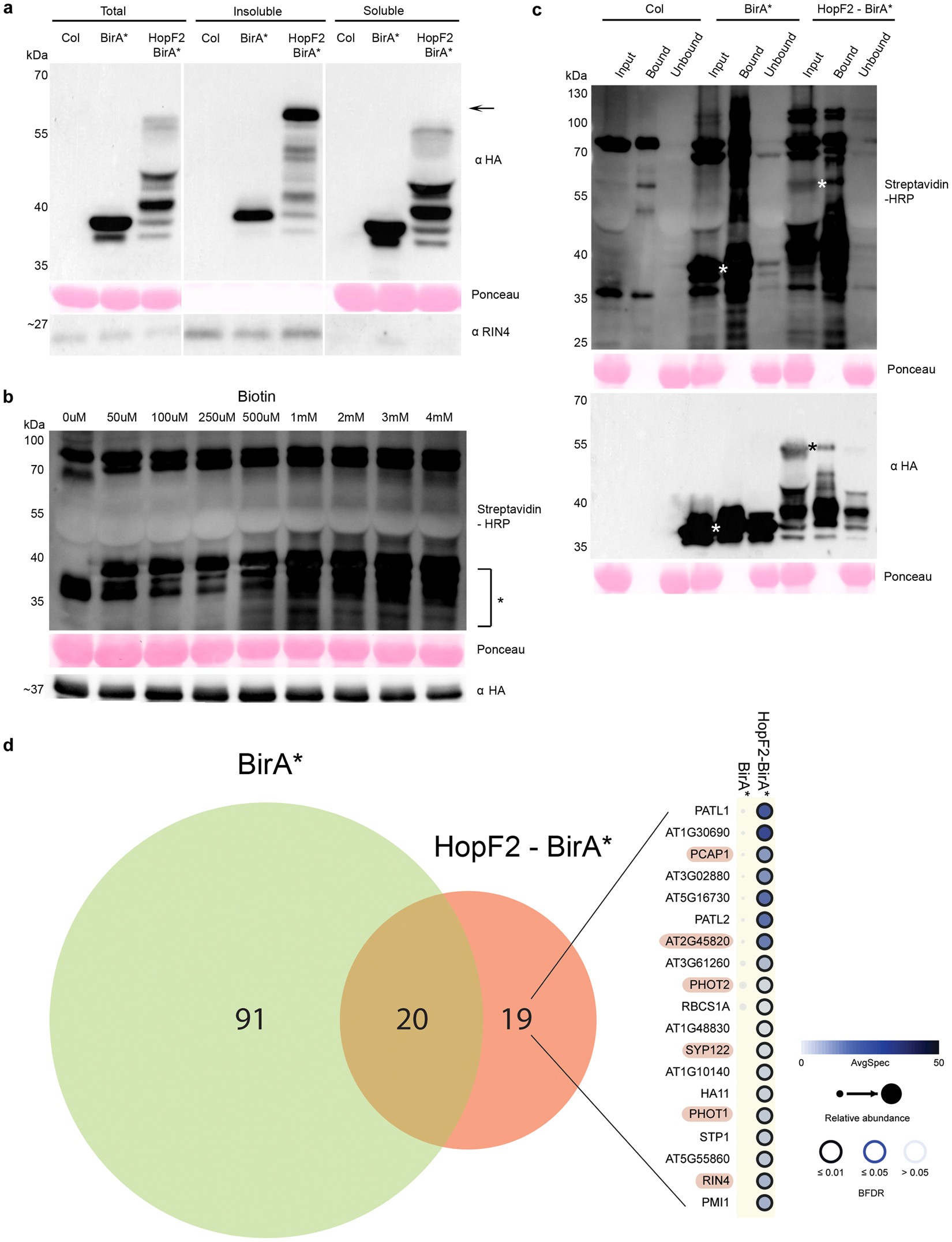
Frontiers | Development of a Rapid in planta BioID System as a Probe for Plasma Membrane-Associated Immunity Proteins

Cells | Free Full-Text | Comparative Application of BioID and TurboID for Protein-Proximity Biotinylation
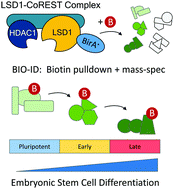
Proximity-dependent biotin identification (BioID) reveals a dynamic LSD1–CoREST interactome during embryonic stem cell differentiation - Molecular Omics (RSC Publishing)
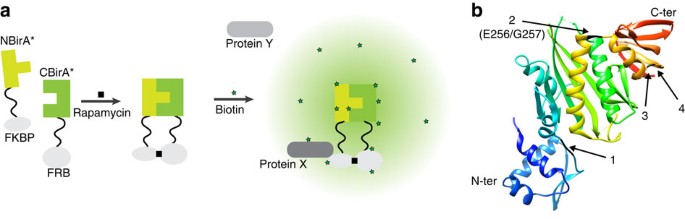
Split-BioID a conditional proteomics approach to monitor the composition of spatiotemporally defined protein complexes | Nature Communications
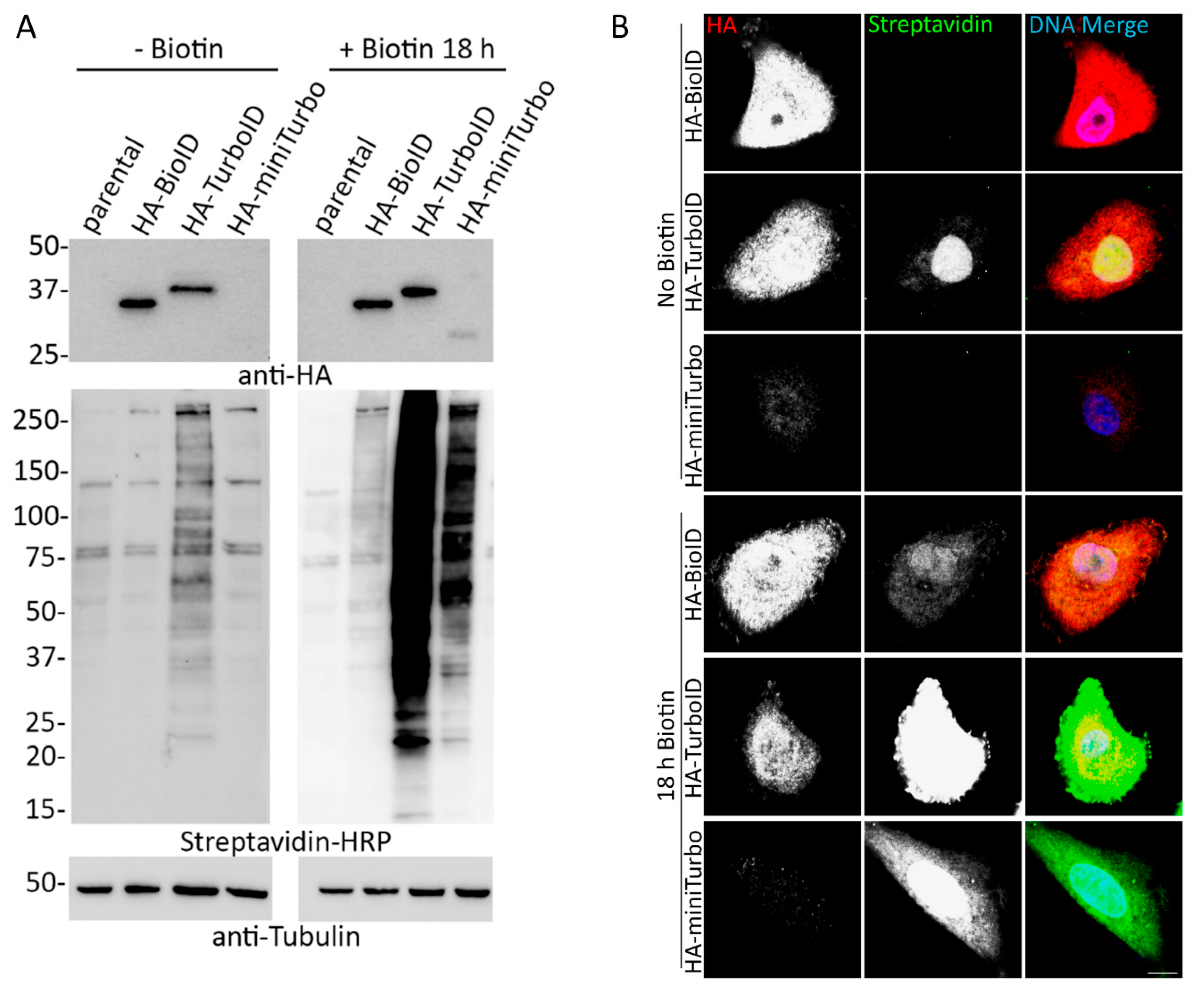
Cells | Free Full-Text | Comparative Application of BioID and TurboID for Protein-Proximity Biotinylation
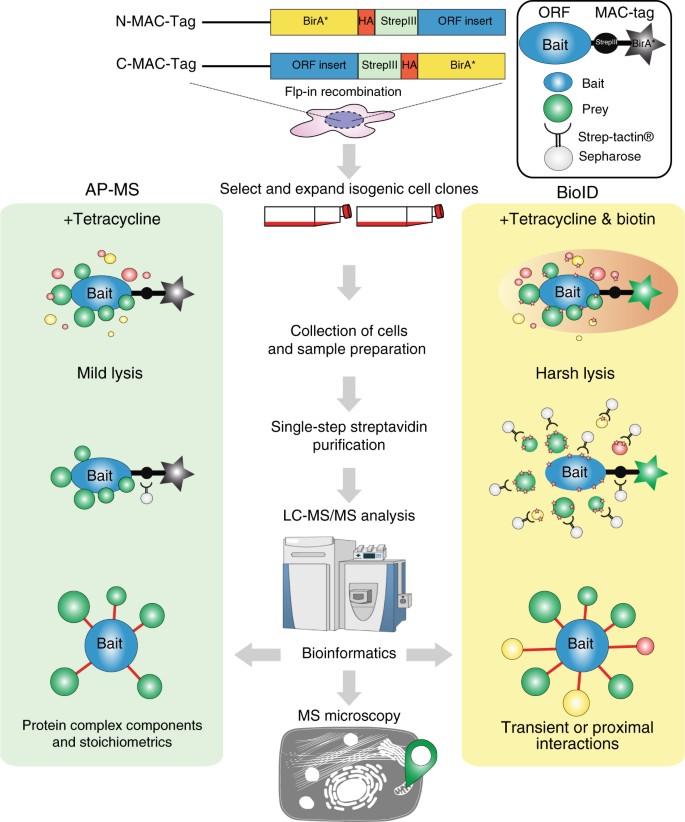
An AP-MS- and BioID-compatible MAC-tag enables comprehensive mapping of protein interactions and subcellular localizations | Nature Communications

Split‐BioID: a proximity biotinylation assay for dimerization‐dependent protein interactions - De Munter - 2017 - FEBS Letters - Wiley Online Library
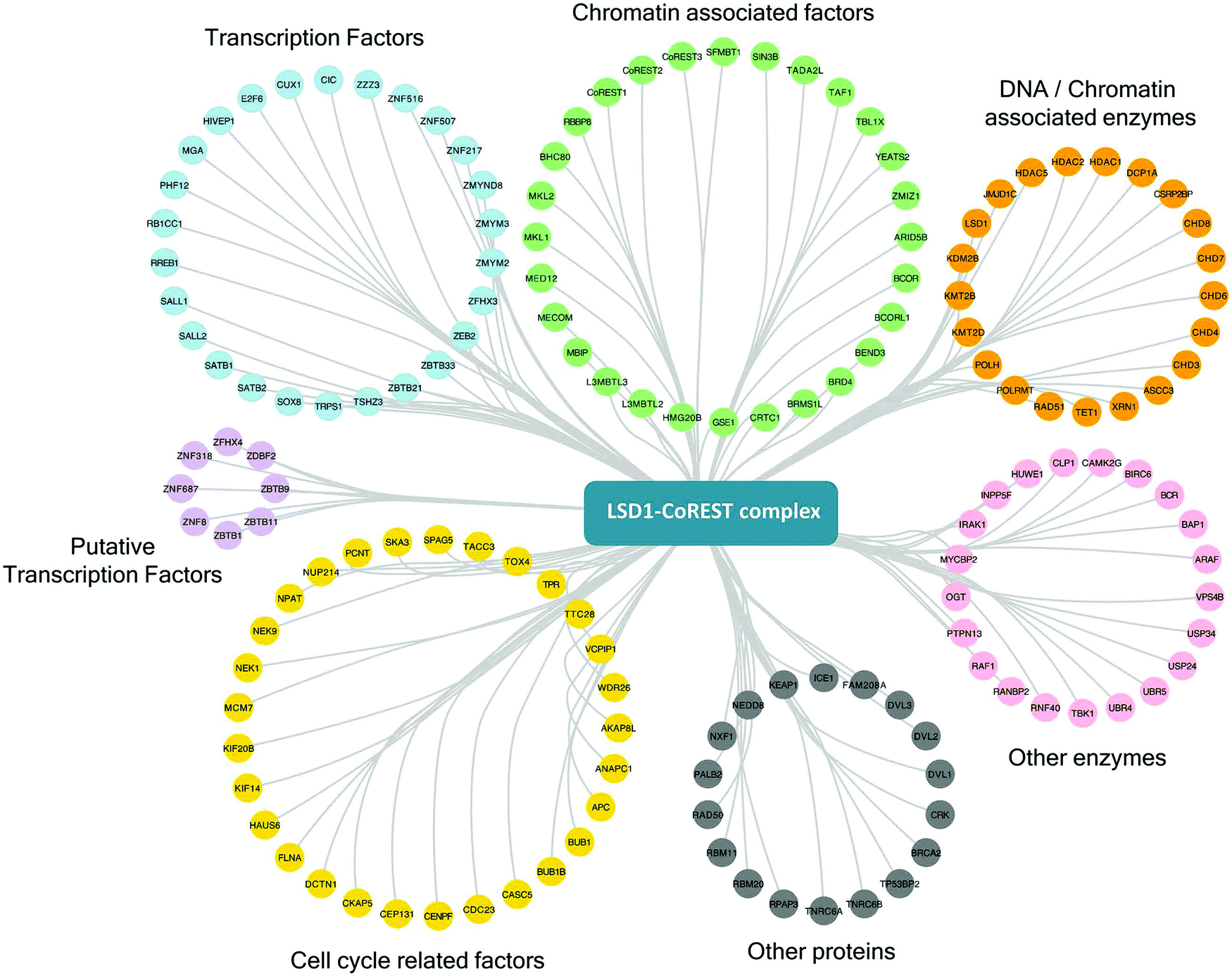
Proximity-dependent biotin identification (BioID) reveals a dynamic LSD1–CoREST interactome during embryonic stem cell differentiation - Molecular Omics (RSC Publishing) DOI:10.1039/D1MO00236H